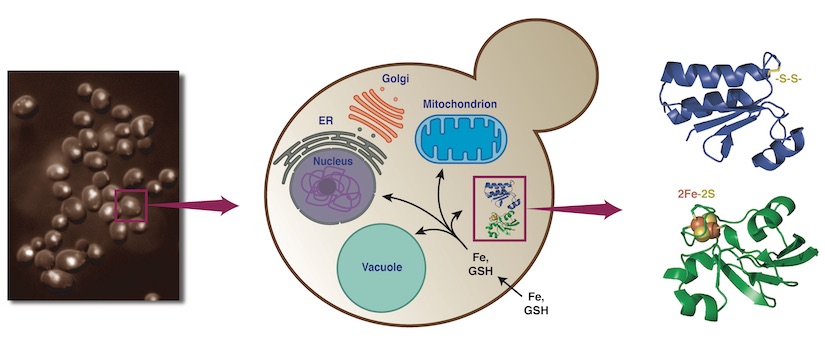
We investigate the molecular mechanisms that coordinate iron metabolism with cellular redox balance, with a particular emphasis on iron-sulfur (Fe-S) cluster biology, glutaredoxins, and metal-dependent regulatory pathways. These systems are highly conserved, yet uniquely adapted in pathogenic fungi to survive in the iron-limited and stressful environment of the host. By defining these mechanisms at the molecular level, our goal is to identify new vulnerabilities in fungal pathogens that can be targeted for antifungal therapy.
Current Research Projects
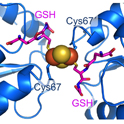
Iron Sensing and Regulatory Circuits
A central question in our lab is: how do cells know when they have enough iron? Our work has helped define how Fe-S clusters function as signaling molecules that control iron-responsive transcription factors in yeast and other fungi. In recent studies, we demonstrated that monothiol glutaredoxins and BolA-like proteins regulate transcriptional repressors through Fe-S cluster transfer and remodeling mechanisms.
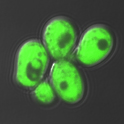
Glutathione and Redox-Dependent Pathways
Glutathione serves as an essential cofactor for anti-oxidant enzymes and plays a key role in metal metabolism. Our lab has a long-standing interest in how glutathione protects cells from oxidative stress and supports Fe-S cluster assembly and transfer. We use genetically-encoded sensors and biochemical approaches to monitor glutathione dynamics in living cells and determine how redox state influences iron-dependent processes.